|
To remove results that come from uncultured, unclassified, or environmental samples you can use all NOT uncultured NOT “environmental” NOT unclassified.įor further examples of Entrez query terms, see here. For example, to only return results from mouse, enter Mus musculus in the Entrez query field. It is possible to restrict the results of a BLAST search to a specific species, or to exclude certain types of sequence by using an Entrez query in the advanced options. For detailed information about these parameters, see the NCBI BLAST Help. 
Changing the scoring table, word size and gap costs will also affect the number of hits returned and the aligned regions. E values higher than this may indicate that the match has occurred by chance. To ensure the results returned include only significant matches, the E value should be set on 1e-3 or less. This is the expanded dialog for Megablast searches: This will bring up program specific settings to allow you to further optimize your search. Advanced Optionsĭepending on the search program selected, you can change the search options by clicking on More Options. When batch searching, all sequences will be compared using the same program against the same database. tblastn – compares protein query to all 6 frames of a translated nucleotide database.blastp – compares protein query to a protein database.blastx – translates the query to protein and searches an amino acid database.Best option for more distantly related query species. blastn – slower but most sensitive option, allows more dissimilar matches.Discontiguous megablast – more sensitive, allows more dissimilar matches, and can be set to ignore certain types of bases.Megablast – fast, but only returns highly similar matches.The following algorithms are available: DNA query: Your display should look something like this:īy default, Geneious Prime will offer a BLAST algorithm most appropriate for the type of query sequence and type of database selected. Click on the Query Centric View tab at the top of the Hit table, then turn off the annotations in the Annotations and Tracks tab, and in the Display tab choose to highlight Disagreements to Reference. 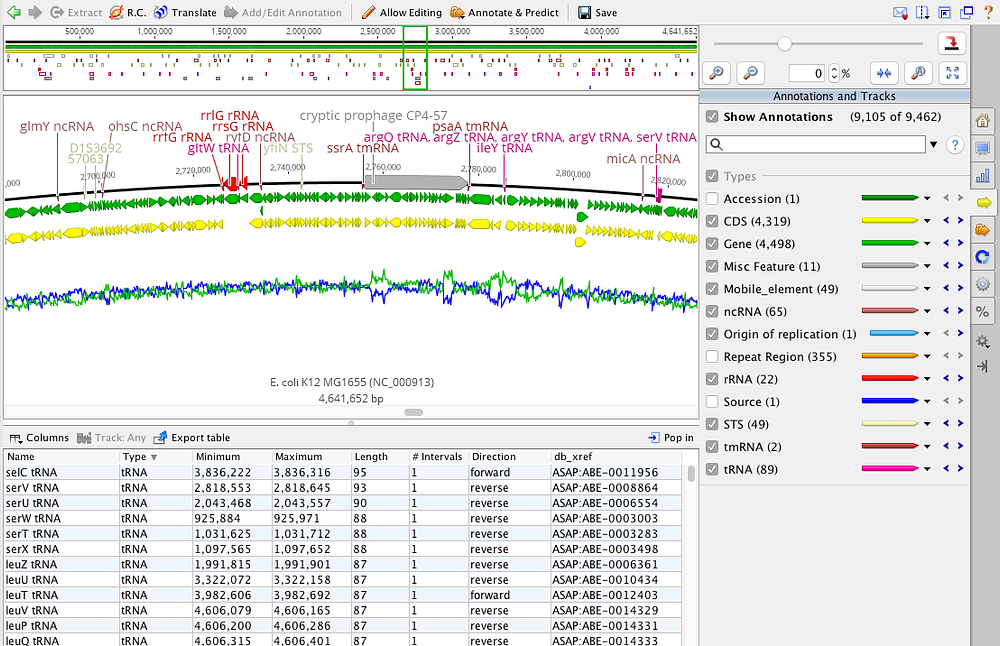
Query-centric view is useful for visualizing all the hits against your query in one window, allowing you to see where conserved regions of your sequence are. This displays the full, annotated sequence of the BLAST hit, with a new “BLAST Hit” annotation showing which region of the sequence matches the query. Once the full sequence is downloaded you’ll see that a Sequence View tab is added to the viewer. To get the full sequence and annotations for the blast hit, click Download Full Sequence(s). The blast hit document returned is a summary document and does not contain the full GenBank record for that sequence. This alignment view only shows the region of alignment between the query and the hit sequence. Like any other alignment in Geneious, you can zoom into display the bases, change the color settings, and highlight agreements or disagreements to the consensus in the General controls to the right of the viewer. 
You can see from the green identity graph above the alignment that the two sequences are identical. Click on the hit to NP_001014408 and you should see something like this: Now that you have a set of search results, you should look at some alignments. Geneious also produces a Grade score, which combines query coverage, e-value and identity values for each hit with weights 0.5, 0.25 and 0.25 respectively, allowing you to determine the longest, highest identity hits. This is why alignments tend to be ranked by their E Value rather than identity. The identity refers only to the aligned region so it is possible to have very short alignments which have high identity. This is because the alignment produced is a local similarity alignment and it has aligned the maximum region it could find between the two sequences. You can see that many of the hits in this example are 100% identical to the query over the length of the alignment, but have different Sequence Lengths. This is also useful as it will indicate how similar the sequence found in the database is to the one you used as a query. In addition to the E Value, there is also a column labelled % Pairwise Identity. You should take these statistics as a guide as there can still be interesting alignments that appear far less significant. 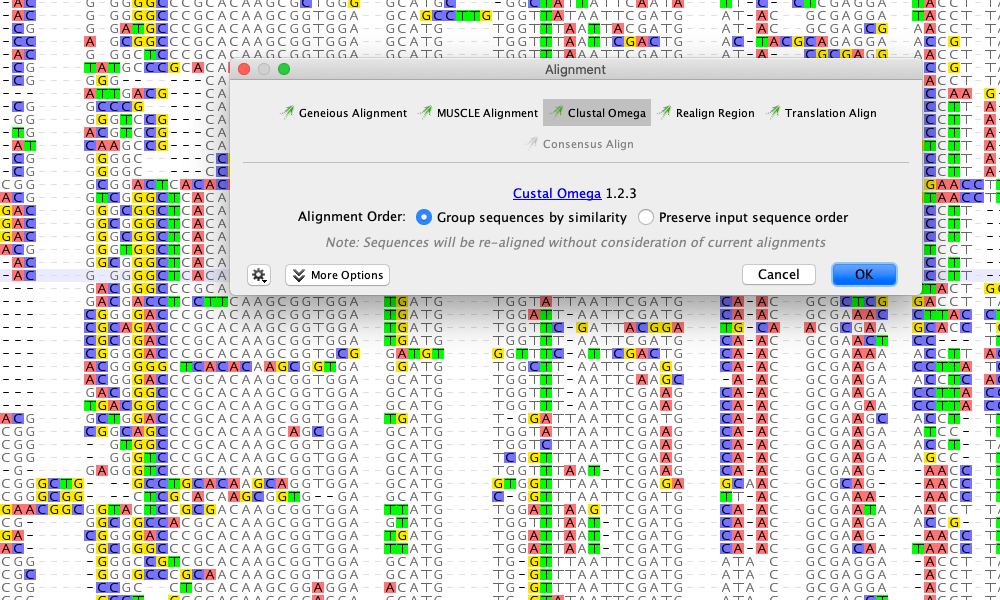
You may even have examples where the E Value reads 0.00e+00 and this is telling you that statistically there is no likelihood that this alignment has happened by chance. This is a very small number and indicates that it is highly unlikely that this alignment would ever occur by chance. The top hit shown here as 1.18e-107 is the same as 1.18×10 -107. For E values, the smaller the number the better.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
